We are at Courant Institute, 251 Mercer St, New York
Computational modeling of cytoskeletal dynamics, cell motility, mitosis, organelle positioning and scaling, galvanotaxis
Ours is a 'dry' lab. We use mathematical and computer modeling, theoretical biophysics and data science tools to develop and simulate models of cell biological processes. We collaborate extensively with experimentalists. Current collaborators are:Keren lab (Technion), Khodjakov lab (Wadsworth Center), Blanchoin/Thery labs in Grenoble and Paris, Baylies lab (Sloan Kettering), Christiaen lab (NYU and U of Bergen), Danuser lab (UT Southwestern). Min Zhao lab (UC Davis), Bershadsky lab (Nat Univ Singapore), Maiato lab (U of Porto), Shvartsman lab (Flatiron Inst and Princeton).
Largely speaking, we are interested in understanding molecular mechanisms of regulation
and mechanics of cell migration, mitosis and cell division and self-organization and
assembly
principles of actin-microtubule-motor machines. We also work on 1) galvanotaxis, and
2) positioning and scaling of multiple nuclei in muscle cells.
Our major current projects are:
1. Detailed quantitative understanding of actin and actomyosin dynamics
(with Danuser, Keren and Blanchoin/Thery labs).
2. Self-assembly, error correction and mechanics of mitotic spindle (with
Khodjakov and Maiato labs).
3. Mechanisms of galvanotaxis (with Zhao lab).
4. Mechanisms of contraction in actin-myosin networks in vitro and in yeast cells (with
Blanchoin/Thery and Keren labs).
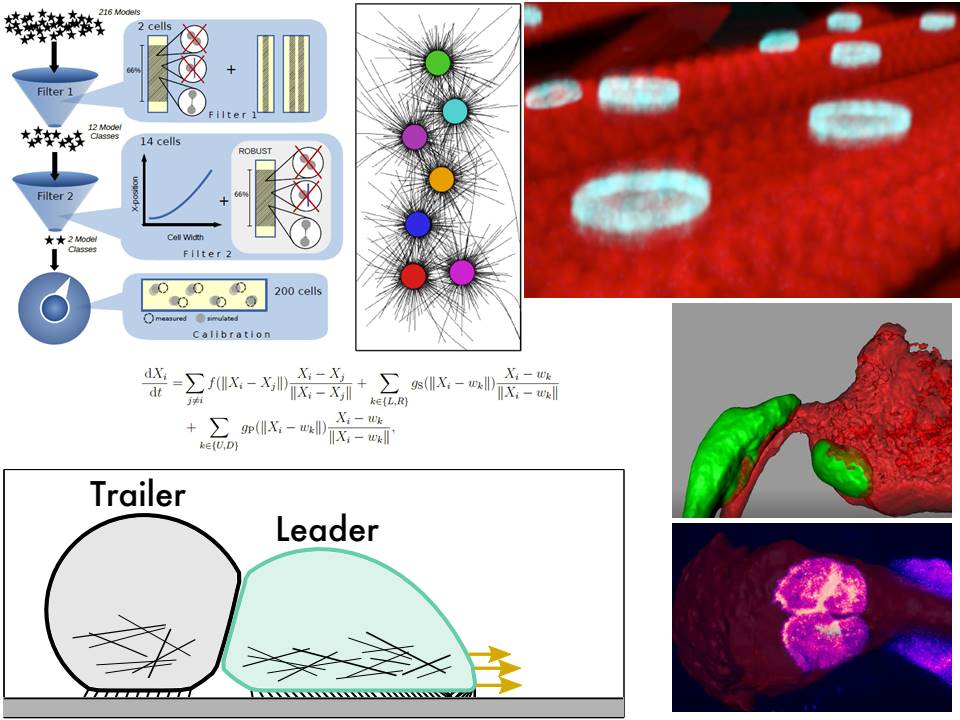
5. Positioning and size scaling of nuclei in multinucleated cells (with Baylies lab).
6. Collective cell migration (with Christiaen lab).
7. Cellular chirality (with Bershadsky lab).
Our publications can be downloaded from here
Who We Are:

What We Do